基本概念
- 外显子测序:外显子测序是指利用序列捕获技术将全基因组外显子区域DNA捕捉并富集后进行高通量测序的基因组分析方法。是一种选择基因组的编码序列的高效策略,外显子测序相对于基因组重测序成本较低,对研究已知基因的SNP、Indel等具有较大的优势。
- GATK(The Genome Analysis Toolkit):是Broad Institute开发的用于二代重测序数据分析的一款软件,是基因分析的工具集。
bcftools
deepvariant
freebayes
bgzip压缩后缀tbi
染色体大的用csi后缀的index
samtools faidx 建index
外显子分析流程

图片alt
使用GATK得到许多的 Call SNP和InDel,输出文件为VCF格式
使用ANNOVAR对VCF格式进行注释,选择与疾病相关的 SNP和InDel
千人基因组, dbSNP注释, AVSIFT, cosmic, esp6500si_all, Clinvar
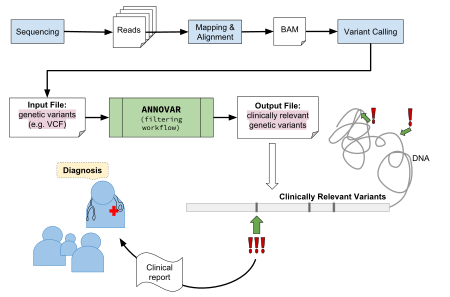
图片alt
https://en.wikipedia.org/wiki/ANNOVAR#Types_of_functional_annotation_of_genetic_variants
突变数据文件
Variant Call Format
##fileformat=VCFv4.3
##fileDate=20090805
##source=myImputationProgramV3.1
##reference=file:///seq/references/1000GenomesPilot-NCBI36.fasta
##contig=<ID=20,length=62435964,assembly=B36,md5=f126cdf8a6e0c7f379d618ff66beb2da,species="Homo sapiens",taxonomy=x>
##phasing=partial
##INFO=<ID=NS,Number=1,Type=Integer,Description="Number of Samples With Data">
##INFO=<ID=DP,Number=1,Type=Integer,Description="Total Depth">
##INFO=<ID=AF,Number=A,Type=Float,Description="Allele Frequency">
##INFO=<ID=AA,Number=1,Type=String,Description="Ancestral Allele">
##INFO=<ID=DB,Number=0,Type=Flag,Description="dbSNP membership, build 129">
##INFO=<ID=H2,Number=0,Type=Flag,Description="HapMap2 membership">
##FILTER=<ID=q10,Description="Quality below 10">
##FILTER=<ID=s50,Description="Less than 50% of samples have data">
##FORMAT=<ID=GT,Number=1,Type=String,Description="Genotype">
##FORMAT=<ID=GQ,Number=1,Type=Integer,Description="Genotype Quality">
##FORMAT=<ID=DP,Number=1,Type=Integer,Description="Read Depth">
##FORMAT=<ID=HQ,Number=2,Type=Integer,Description="Haplotype Quality">
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT NA00001 NA00002 NA00003
20 14370 rs6054257 G A 29 PASS NS=3;DP=14;AF=0.5;DB;H2 GT:GQ:DP:HQ 0|0:48:1:51,51 1|0:48:8:51,51 1/1:43:5:.,.
20 17330 . T A 3 q10 NS=3;DP=11;AF=0.017 GT:GQ:DP:HQ 0|0:49:3:58,50 0|1:3:5:65,3 0/0:41:3
20 1110696 rs6040355 A G,T 67 PASS NS=2;DP=10;AF=0.333,0.667;AA=T;DB GT:GQ:DP:HQ 1|2:21:6:23,27 2|1:2:0:18,2 2/2:35:4
20 1230237 . T . 47 PASS NS=3;DP=13;AA=T GT:GQ:DP:HQ 0|0:54:7:56,60 0|0:48:4:51,51 0/0:61:2
20 1234567 microsat1 GTC G,GTCT 50 PASS NS=3;DP=9;AA=G GT:GQ:DP 0/1:35:4 0/2:17:2 1/1:40:3
MAF Format
下游分析
- 已知基因,了解基因在各个癌症中的突变情况
- 已知肿瘤,了解肿瘤中那些基因突变频率高
- 已知肿瘤,了解肿瘤中那些位点突变频率高
- 按临床分组,了解不同组之间突变频率